CUT&Tag vs CUT&RUN Which Epigenomics Method Should You Choose?
CUT&Tag vs CUT&RUN: Which Epigenomics Method Should You Choose?
The field of epigenetics has undergone a major transformation with the development of advanced chromatin profiling techniques. Among these, CUT&Tag and CUT&RUN have emerged as powerful alternatives to traditional methods like ChIP-seq.
Both approaches offer high sensitivity, low background noise, and reduced sample requirements, but they differ in mechanism, workflow, and optimal applications. This article provides a deep scientific comparison to help researchers select the most appropriate method.
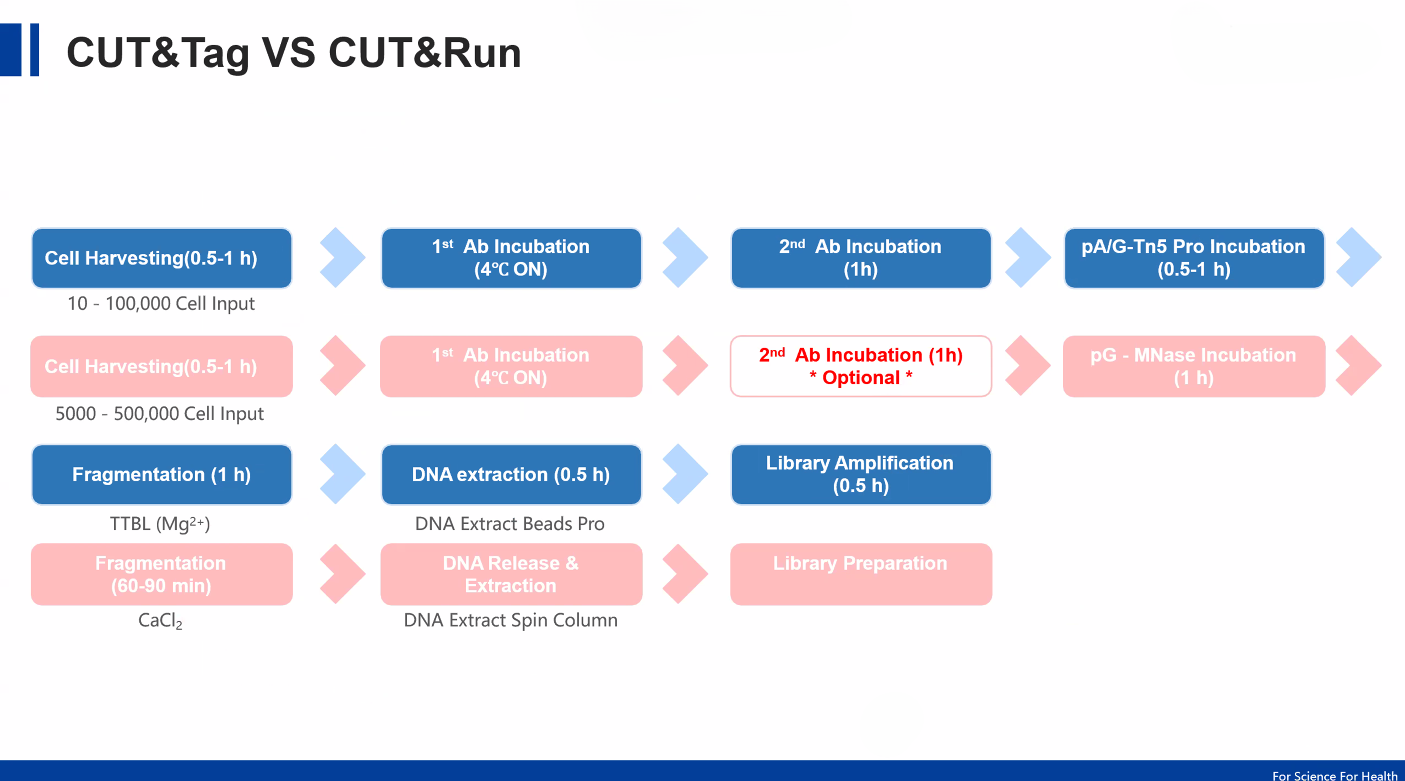
Understanding the Core Principles
CUT&RUN Mechanism
CUT&RUN relies on antibody-targeted cleavage using a Protein A–MNase (pA-MNase) fusion enzyme.
Key Steps:
-
Antibody binds to target protein or histone mark
-
pA-MNase is recruited to the binding site
-
MNase cleaves DNA near the target
-
Released DNA fragments diffuse into solution
-
DNA is purified and prepared for sequencing
This method generates precise fragments with minimal background noise
CUT&Tag Mechanism
CUT&Tag builds upon CUT&RUN but introduces direct tagmentation using a Protein A/G–Tn5 transposase (pAG-Tn5).
Key Steps:
-
Antibody binds to target chromatin feature
-
pAG-Tn5 is recruited
-
Tn5 inserts sequencing adapters directly into DNA
-
DNA fragments are PCR-amplified without library prep
This enables a faster and more streamlined workflow
CUT&Tag vs CUT&RUN: Head-to-Head Comparison
1. Cell Input Requirements
CUT&RUN: ~10⁴–10⁵ cells
CUT&Tag: ≤10³ cells (even single-cell)
CUT&Tag (ultra-low input capability)
2. Background Noise
-
CUT&RUN: Very low
-
CUT&Tag: Extremely low
CUT&Tag (higher signal-to-noise ratio)
3. Workflow Complexity
CUT&RUN: Requires DNA extraction + library preparation
CUT&Tag: Direct PCR (no separate library prep)
CUT&Tag (simpler workflow)
4. Cost Efficiency
CUT&RUN: Moderate (~$100–300/sample)
CUT&Tag: Moderate to low (~$100–300/sample, scalable)
Tie, but CUT&Tag often reduces sequencing depth further
5. Resolution and Sensitivity
CUT&RUN: High resolution
CUT&Tag: Ultra-high resolution
CUT&Tag
6. Fragmentation Strategy
CUT&RUN: Enzymatic cleavage via MNase
CUT&Tag: Tagmentation via Tn5 (cuts + tags DNA)
Fundamental difference impacting downstream workflow
Summary Table
| Feature | CUT&RUN | CUT&Tag |
|---|---|---|
| Enzyme | pA-MNase | pAG-Tn5 |
| Input Requirement | Low | Very low / single-cell |
| Background Noise | Low | Very low |
| Library Prep | Required | Not required |
| Resolution | High | Very high |
| Workflow Time | ~2 days | ~1.5–2 days |
Applications: When to Use Each Method?
? Choose CUT&RUN if:
You want a well-established protocol
You are working with moderate sample input
You prefer controlled DNA fragmentation
? Choose CUT&Tag if:
You have limited or rare samples
You need single-cell epigenomics
You want a faster, simplified workflow
You aim for maximum sensitivity
Commercial Kits and Optimization
Many biotechnology companies, including Vazyme, offer optimized kits for both CUT&RUN and CUT&Tag.
Features of Modern Kits:
Pre-loaded enzymes (pA-MNase or pAG-Tn5)
Optimized buffers for low background
Indexed primers for multiplex sequencing
Protocols compatible with low-input samples
These kits significantly enhance reproducibility and reduce hands-on time.
Experimental Considerations
To ensure success in both methods:
✔ Antibody Specificity
Critical for both CUT&RUN and CUT&Tag
Poor specificity leads to false peaks
✔ Enzyme Optimization
Over-digestion (CUT&RUN) or over-tagmentation (CUT&Tag) can increase noise
✔ Sample Quality
Intact nuclei are essential for accurate profiling
Future of Epigenomics
The transition from ChIP-seq to CUT&RUN and CUT&Tag reflects a broader shift toward:
Low-input technologies
Single-cell analysis
Cost-efficient sequencing
Among the two, CUT&Tag is rapidly becoming the preferred method due to its scalability and simplicity.
Conclusion
Both CUT&RUN and CUT&Tag represent major advancements in epigenetic profiling, offering significant improvements over traditional methods. While CUT&RUN provides a reliable and precise approach, CUT&Tag pushes the boundaries with ultra-low input requirements, simplified workflow, and superior resolution.
For most modern applications, especially in single-cell and low-input studies, CUT&Tag is the method of choice.
Recent Posts
-
Herpes Simplex Virus 2 (HSV-2)
Herpes Simplex Virus 2 (HSV-2): A Comprehensive Guide Introduction Herpes Simplex Virus 2 (HSV-2) is …2nd Apr 2026 -
Homologous Chromosomes
Homologous Chromosomes: Key Traits & Biological Significance Introduction Homologous chromosomes are …2nd Apr 2026 -
CUT&Tag vs CUT&RUN Which Epigenomics Method Should You Choose?
CUT&Tag vs CUT&RUN: Which Epigenomics Method Should You Choose? The field of epigenetics has undergo …17th Mar 2026