CUT&Tag A Next-Generation Method for High-Resolution Epigenomic Profiling
CUT&Tag A Next-Generation Method for High-Resolution Epigenomic Profiling
The rapid evolution of epigenomics has driven the need for more sensitive, efficient, and cost-effective methods to study chromatin dynamics. Among the most advanced techniques available today, CUT&Tag has emerged as a powerful alternative to traditional chromatin immunoprecipitation approaches.
This method enables precise mapping of protein–DNA interactions, histone modifications, and transcription factor binding with ultra-low input requirements, making it particularly valuable in modern molecular biology and single-cell research.
What is CUT&Tag?
CUT&Tag (Cleavage Under Targets and Tagmentation) is an epigenomic profiling technique that combines antibody-based targeting with Tn5 transposase-mediated tagmentation. Unlike conventional methods such as ChIP-seq, CUT&Tag integrates DNA fragmentation and adapter ligation into a single step, significantly simplifying the workflow.
The method relies on a fusion protein (Protein A/G–Tn5 transposase) that binds to antibodies targeting specific chromatin features, allowing direct insertion of sequencing adapters at precise genomic locations.
Molecular Mechanism of CUT&Tag
The CUT&Tag workflow is built on a highly targeted enzymatic process:
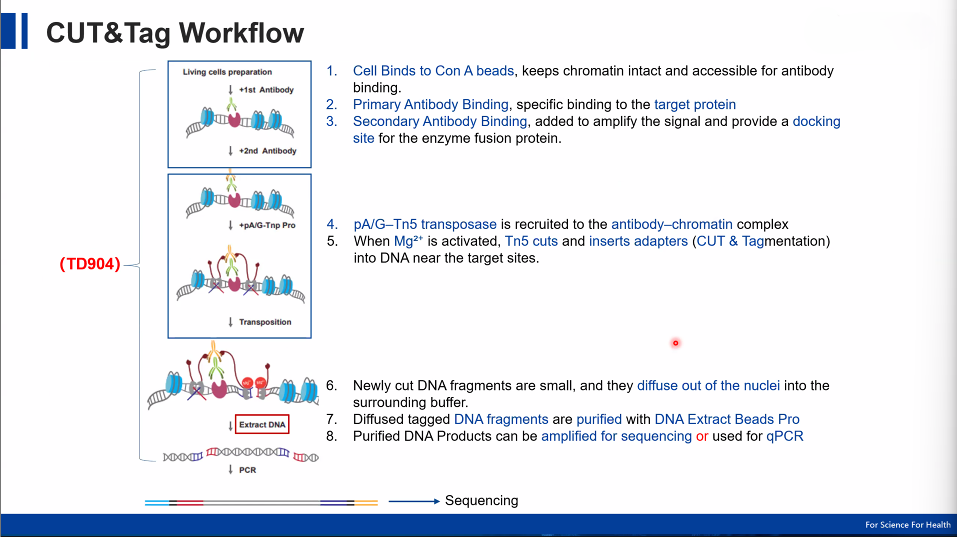
1. Antibody Binding
Cells or nuclei are immobilized and incubated with a primary antibody specific to:
Histone modifications (e.g., H3K27ac, H3K4me3)
Transcription factors
Chromatin-associated proteins
2. Secondary Antibody Amplification
A secondary antibody enhances signal by increasing binding sites for the transposase complex.
3. pA/G-Tn5 Recruitment
A fusion protein combining Protein A/G and Tn5 transposase binds to the antibody–chromatin complex.
4. Targeted Tagmentation
Upon activation, Tn5 simultaneously:
Cleaves DNA near the binding site
Inserts sequencing adapters
This step eliminates the need for:
Mechanical fragmentation (e.g., sonication)
Separate adapter ligation
5. PCR Amplification
Adapter-ligated DNA fragments are directly amplified via PCR, producing libraries ready for sequencing.
CUT&Tag Workflow Overview
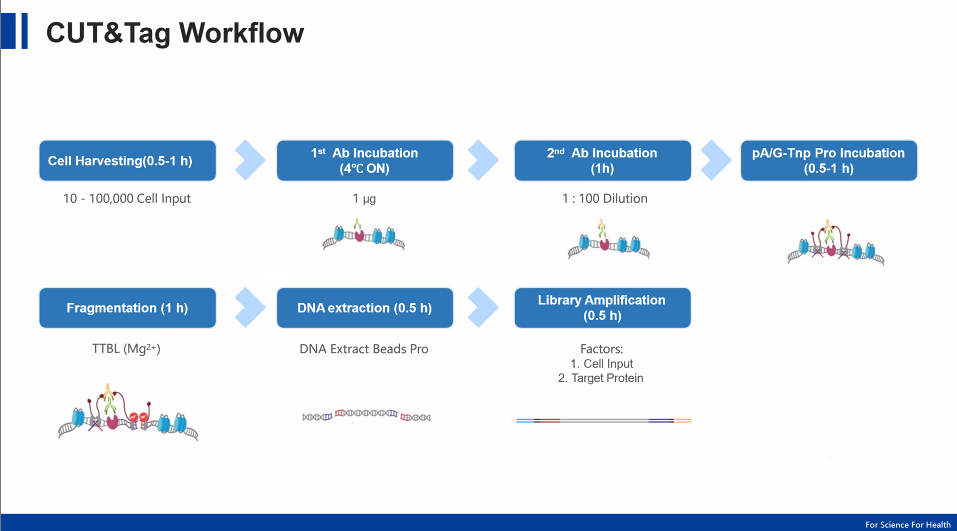
Here is a simplified experimental workflow:
-
Cell or nuclei preparation
-
Antibody incubation
-
pA/G-Tn5 binding
-
Tagmentation activation
-
DNA release and PCR amplification
-
Next-generation sequencing
Total time: ~1.5–2 days, significantly faster than traditional methods
Advantages of CUT&Tag
? Ultra-Low Input Requirement
CUT&Tag can be performed with:
As few as 1,000 cells
Even single-cell resolution
? High Signal-to-Noise Ratio
Because tagmentation occurs only at antibody-bound sites:
Background noise is extremely low
Data quality is significantly improved
? Reduced Cost
Lower sequencing depth required
No need for extensive library preparation
Typical cost: ~$100–300 per sample
? High Resolution
Precise insertion of adapters enables:
Base-pair-level resolution
Accurate peak detection
? Simplified Workflow
No crosslinking
No sonication
Fewer purification steps
CUT&Tag vs Other Epigenetic Methods
Compared to:
-
ChIP-seq → CUT&Tag is faster, cheaper, and requires fewer cells
-
CUT&RUN → CUT&Tag offers higher resolution and simpler library prep
CUT&Tag is currently considered one of the most efficient epigenomic profiling techniques available
Applications of CUT&Tag
CUT&Tag is widely used in:
Histone Modification Mapping
H3K27ac (active enhancers)
H3K4me3 (promoters)
H3K27me3 (repressive marks)
Transcription Factor Binding Analysis
Identification of regulatory elements
Gene expression control studies
Cancer Epigenetics
Tumor-specific chromatin signatures
Biomarker discovery
Single-Cell Epigenomics
Cellular heterogeneity analysis
Developmental biology studies

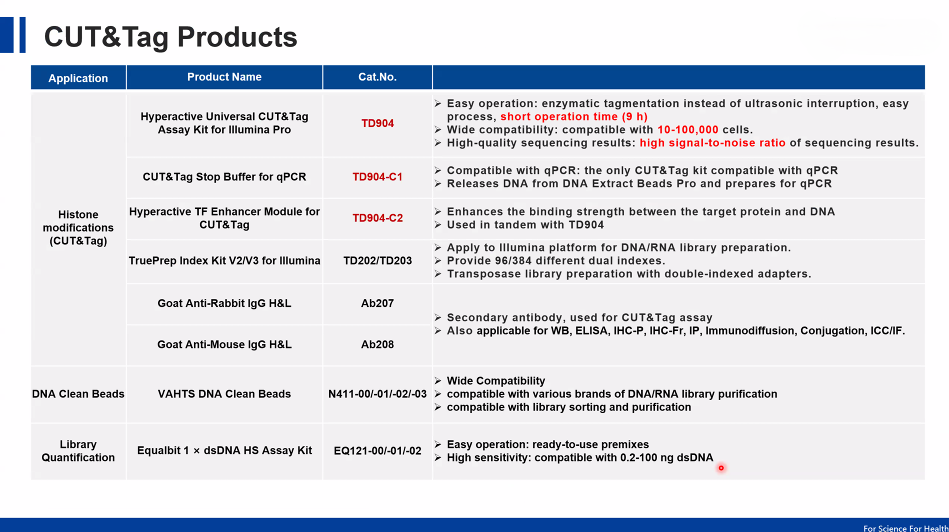
CUT&Tag Kits and Reagents (Product Perspective)
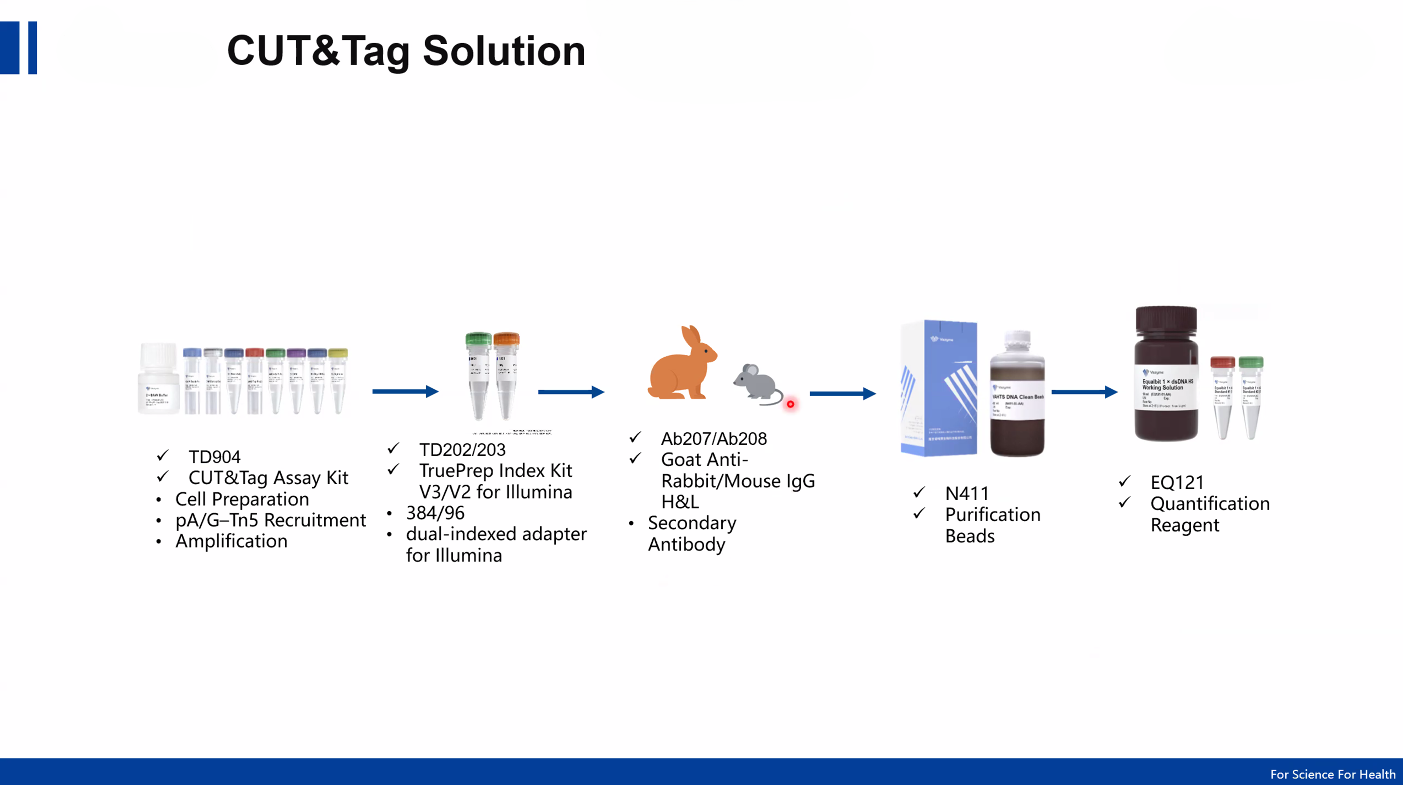
Modern CUT&Tag workflows are supported by optimized commercial kits which include:
Pre-loaded pA/G-Tn5 transposase
High-specificity buffers
Optimized reaction conditions
Indexed primers for multiplex sequencing
Key Features of Advanced Kits:
Streamlined protocols (≤2 days)
Minimal hands-on time
High reproducibility
Compatibility with low-input samples
These kits significantly reduce experimental variability and make CUT&Tag accessible to a broader range of laboratories.
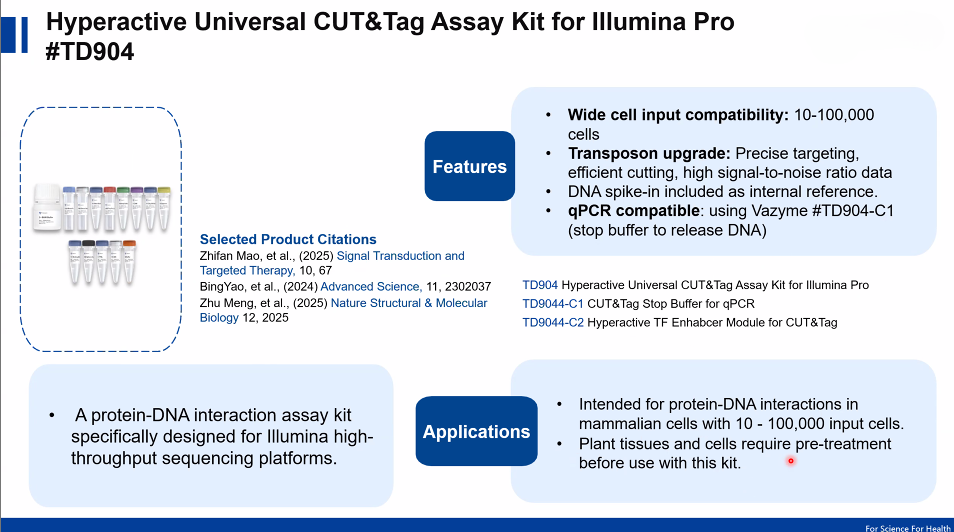

Experimental Considerations
To achieve optimal CUT&Tag results:
✔ Antibody Quality
Use highly specific, validated antibodies
Poor antibody quality = weak signal
✔ Cell Permeabilization
Proper permeabilization ensures enzyme access
Over-treatment can damage chromatin
✔ Enzyme Titration
Excess Tn5 may increase background
Optimization is key
✔ Sequencing Depth
Lower than ChIP-seq, but still critical for rare targets
Data Analysis Pipeline
CUT&Tag data analysis typically involves:
-
Quality control (FastQC)
-
Read alignment (Bowtie2/BWA)
-
Peak calling (MACS2)
-
Visualization (IGV, genome browsers)
Due to low background, peak detection is more precise and reliable
Future Perspectives
CUT&Tag is shaping the future of epigenomics by enabling:
Integration with multi-omics approaches
High-throughput single-cell profiling
Clinical applications in precision medicine
As sequencing technologies continue to evolve, CUT&Tag is expected to become a standard tool in epigenetic research and diagnostics.
Conclusion
CUT&Tag represents a breakthrough in chromatin profiling, offering unmatched sensitivity, resolution, and efficiency. Its ability to generate high-quality data from minimal input material makes it an essential tool for modern epigenetics research.
For laboratories seeking to optimize performance while reducing cost and complexity, CUT&Tag is rapidly becoming the method of choice.
Recent Posts
-
Herpes Simplex Virus 2 (HSV-2)
Herpes Simplex Virus 2 (HSV-2): A Comprehensive Guide Introduction Herpes Simplex Virus 2 (HSV-2) is …2nd Apr 2026 -
Homologous Chromosomes
Homologous Chromosomes: Key Traits & Biological Significance Introduction Homologous chromosomes are …2nd Apr 2026 -
CUT&Tag vs CUT&RUN Which Epigenomics Method Should You Choose?
CUT&Tag vs CUT&RUN: Which Epigenomics Method Should You Choose? The field of epigenetics has undergo …17th Mar 2026