ChIP-seq vs CUT&RUN vs CUT&Tag
ChIP-seq vs CUT&RUN vs CUT&Tag: Advanced Methods to Study Epigenetics
Epigenetics research has rapidly evolved with the development of high-resolution sequencing technologies. Among the most widely used approaches, ChIP-seq, CUT&RUN, and CUT&Tag stand out as essential tools for mapping protein–DNA interactions and histone modifications.
This article provides a detailed comparison of these techniques, helping researchers choose the most suitable method depending on experimental needs, cost, and sample availability.
Overview of Epigenetic Profiling Technologies
Epigenetic mechanisms such as histone modifications and transcription factor binding regulate gene expression without altering DNA sequences. Techniques like ChIP-seq, CUT&RUN, and CUT&Tag enable genome-wide mapping of these regulatory events with increasing precision and efficiency.
ChIP-seq: The Traditional Gold Standard
ChIP-seq remains one of the most established methods in epigenomics.
Key Features:
Requires high cell input (≥10⁶ cells)
Uses crosslinking and sonication to fragment DNA
Relies heavily on high-quality antibodies
Produces genome-wide protein–DNA interaction maps
Advantages:
Highly standardized and reproducible
Suitable for broad histone marks and transcription factors
Limitations:
High background noise
Expensive sequencing costs (~$500–1000/sample)
Long protocol duration (~5 days)
Use case: Large-scale studies and legacy dataset comparison
CUT&RUN: Precision with Lower Input
CUT&RUN is a newer method designed to overcome limitations of ChIP-seq.
Key Features:
Requires low cell input (10⁴–10⁵ cells)
Uses pG-MNase enzyme for targeted DNA cleavage
No crosslinking required (native chromatin)
Advantages:
Low background noise
Faster workflow (~2 days)
Reduced sequencing depth required
Limitations:
Requires library preparation
Less standardized than ChIP-seq
CUT&Tag: Next-Generation Epigenomics
CUT&Tag represents a major technological advancement in epigenetics.
Key Features:
Works with ultra-low input (≤10³ cells or single-cell)
Uses pAG-Tn5 transposase for direct adapter insertion
No need for traditional library preparation
Advantages:
Very low background noise
High-resolution mapping
Cost-effective (~$100–300/sample)
Scalable for single-cell epigenomics
Limitations:
Requires careful optimization
Still emerging in some labs
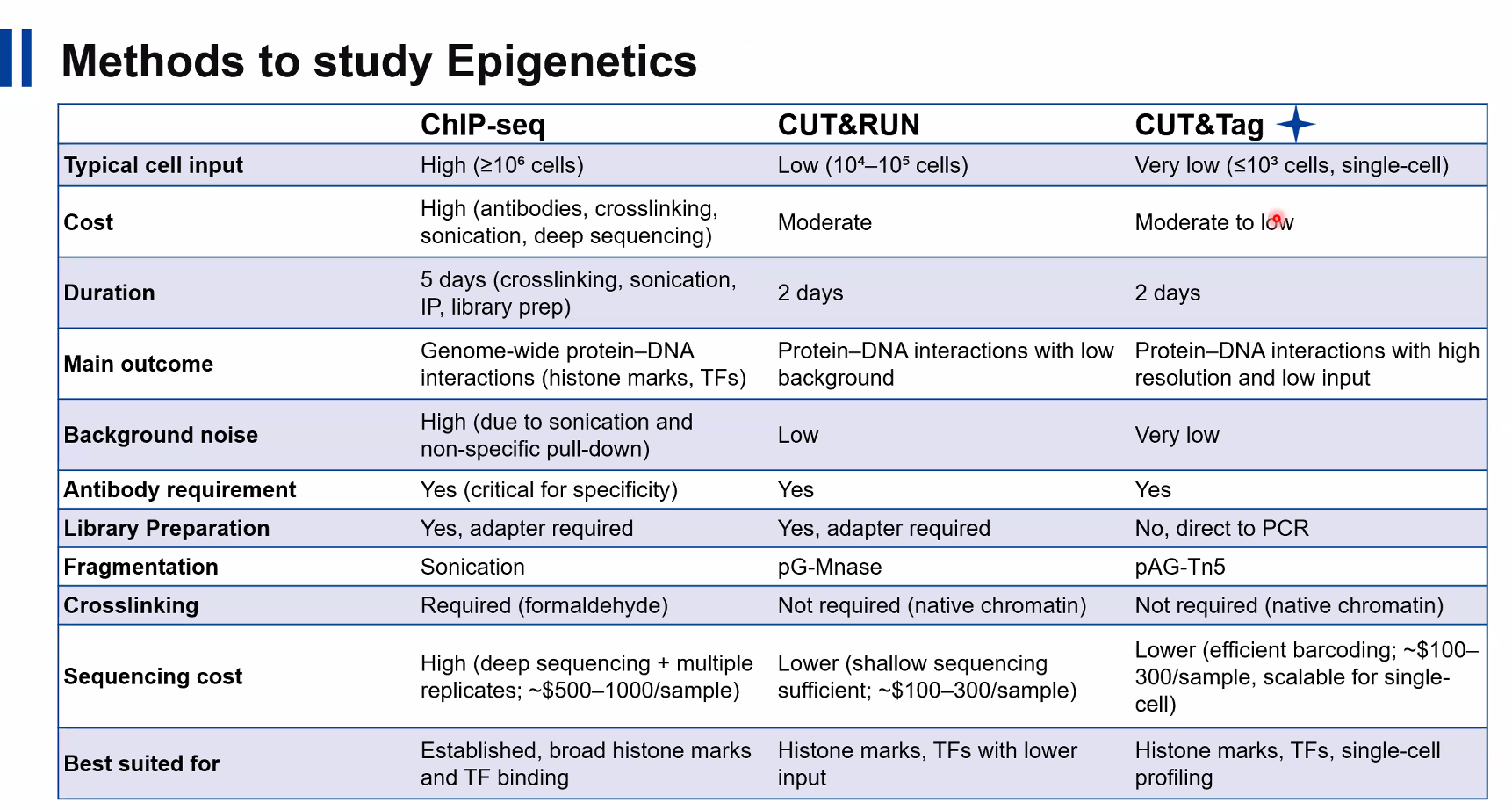
Comparative Analysis of ChIP-seq, CUT&RUN, and CUT&Tag
The image above highlights key differences between these three techniques:
Cell Input:
CUT&Tag < CUT&RUN < ChIP-seq
Background Noise:
CUT&Tag (very low) < CUT&RUN (low) < ChIP-seq (high)
Cost & Efficiency:
CUT&Tag and CUT&RUN are significantly more cost-effective than ChIP-seq
Workflow Time:
CUT&RUN and CUT&Tag (~2 days) are faster than ChIP-seq (~5 days)
Resolution:
CUT&Tag provides the highest resolution and sensitivity
Which Epigenetics Method Should You Choose?
Choosing between these techniques depends on your experimental constraints:
| Scenario | Recommended Method |
|---|---|
| Large sample size, standardized workflows | ChIP-seq |
| Limited cells, high accuracy | CUT&RUN |
| Single-cell or ultra-low input | CUT&Tag |
Future Trends in Epigenomics
The shift from ChIP-seq toward CUT&RUN and CUT&Tag reflects a broader trend in epigenomics:
-
Reduced sample requirements
-
Higher resolution data
-
Lower sequencing costs
-
Integration with single-cell technologies
These advances are enabling deeper insights into gene regulation, cancer biology, and developmental processes.
Conclusion
While ChIP-seq remains a reliable and widely used technique, modern alternatives like CUT&RUN and CUT&Tag offer superior performance in terms of sensitivity, cost, and efficiency. As epigenetics research continues to evolve, these next-generation methods are becoming the preferred choice for high-resolution and low-input applications.
Recent Posts
-
Herpes Simplex Virus 2 (HSV-2)
Herpes Simplex Virus 2 (HSV-2): A Comprehensive Guide Introduction Herpes Simplex Virus 2 (HSV-2) is …2nd Apr 2026 -
Homologous Chromosomes
Homologous Chromosomes: Key Traits & Biological Significance Introduction Homologous chromosomes are …2nd Apr 2026 -
CUT&Tag vs CUT&RUN Which Epigenomics Method Should You Choose?
CUT&Tag vs CUT&RUN: Which Epigenomics Method Should You Choose? The field of epigenetics has undergo …17th Mar 2026